Ribosomes
Machines and more
Classified as a type of molecular machine, ribosomes are universally present in all nucleus-containing cells, where they play a central role in the manufacture of proteins. Using strands of messenger RNA (mRNA) as a template, ribosomes are able to link amino acids together in chains that constitute the primary structure of a new protein.
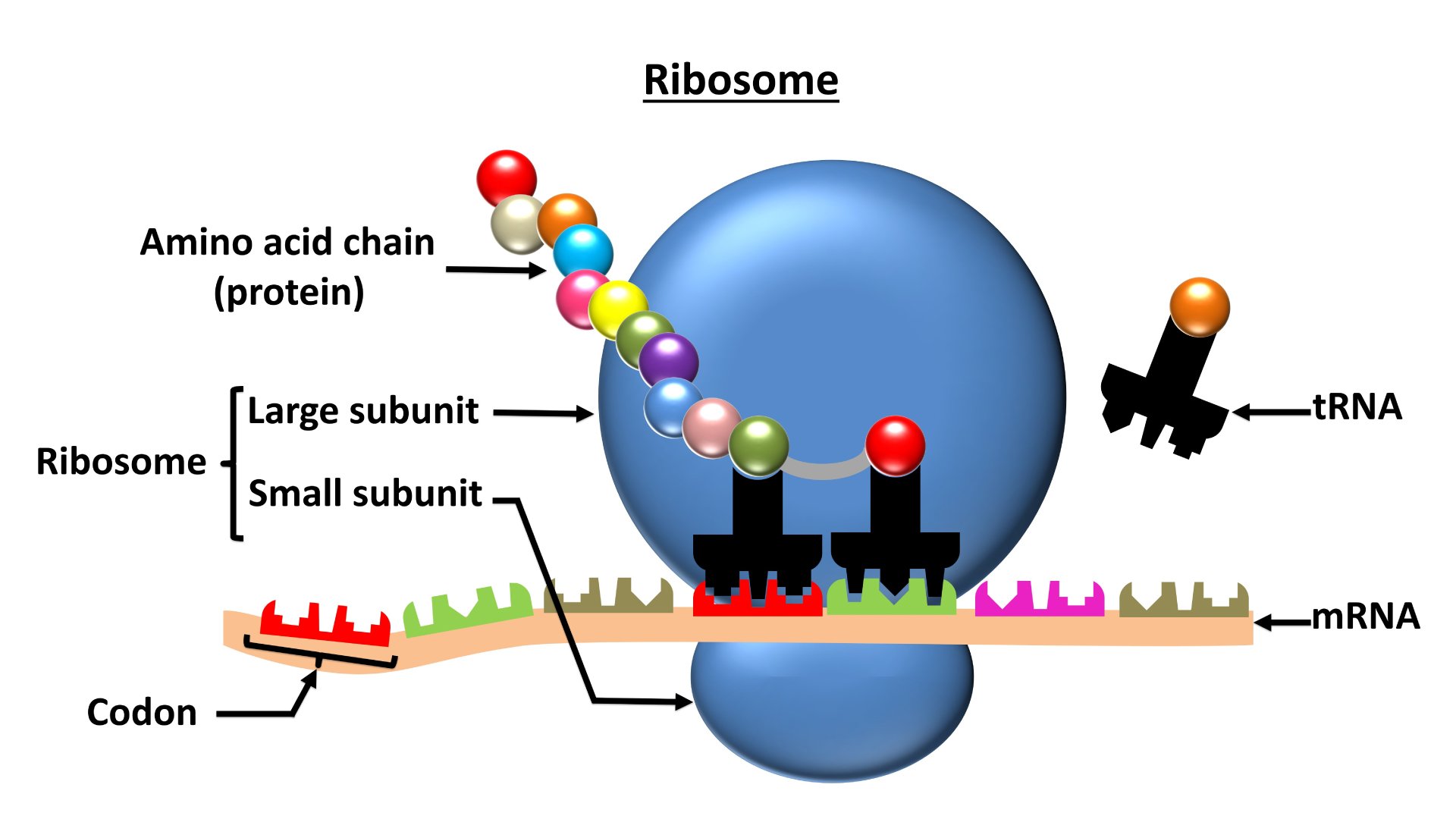
Whether it's bound to an intracellular membrane or "free floating" in cellular cytoplasm, a single ribosome is assembled from two subunits, large and small. The entire structure is described as a "ribonucleoprotein complex," i.e., consisting solely of folded ribosomal RNA (rRNA) and associated proteins.
In prokaryotic cells, the two large and small subunits, termed 50S and 30S, respectively (for their sedimentation coefficients during ultracentrifugal separation), differ not just in size, but in function:
- 30S provides decoding activity and binds to the mRNA template
- 50S functions catalytically and is bound to the aminoacylated ("charged") transfer RNA (tRNA) complexes essential for translation and protein synthesis
Ribosomes in eukaryotic cells are similarly assembled from 40S and 60S subunits intertwined with their own associated proteins. The most biochemically active regions within all ribosomes, however, are remarkably conserved across kingdoms—a reflection of their dependence on structurally universal mRNA and amino acid substrates, as well as evidence for their common evolutionary origin.
The advantages of being different
Significant variations in structure and function between prokaryotic and eukaryotic ribosomes exist, nonetheless. This makes it possible to select antibiotics that specifically target bacterial ribosomes while leaving human ribosomes unharmed. Below are some common ribosome-targeting antibiotics and their mechanisms of action.
| ANTIBIOTIC | ACTION |
| Streptomycin & other aminoglycosides | Inhibit initiation and cause misreading of mRNA (prokaryotes) |
| Tetracycline | Binds to the 30S subunit and inhibits binding of aminoacyl-tRNAs (prokaryotes) |
| Chloramphenicol | Inhibits the peptidyl transferase activity of the 50S ribosomal subunit (prokaryotes) |
| Cycloheximide | Inhibits the peptidyl transferase activity of the 60S ribosomal subunit (eukaryotes; used in vitro for yeast and mold suppression) |
| Erythromycin | Binds to the 50S subunit and inhibits translocation (prokaryotes) |
| Puromycin | Causes premature chain termination by acting as an analog of aminoacyl-tRNA (prokaryotes and eukaryotes) |
Source: https://www.ncbi.nlm.nih.gov/books/NBK22531/
Current topics in ribosome research
More recent discoveries related to ribosome structure and function have opened some promising paths for future study:- Ribosomes as active regulators of gene expression (as opposed to the passive "machines" they've been previously considered)
- Ribosomopathies — undesirable alterations in ribosome structure or biogenesis that result in human disease conditions
- Ribosomes "breaking the rules" of conventional gene translation by:
- Bypassing stop codons to continue translation
- Skipping over several codons before resuming normally
- Programmed frameshifting resulting in alternative gene products
Still other research seeks to experimentally "retro-evolve" ribosomes to a hypothesized simpler, primordial form that may help answer lingering questions about the origin of protein synthesis — a concept closely linked with the origin of life itself.

