T Helper Cells
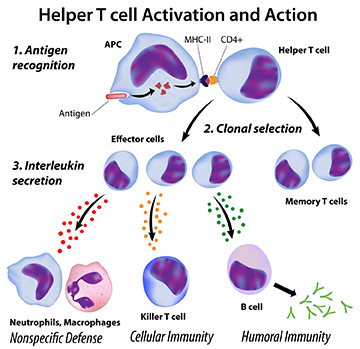
T helper (Th) cells are CD4+ T cells that play a central role in the adaptive immune system, by facilitating the activity of other immune cells – especially antigen-presenting cells (APCs) such as B cells, macrophages, and dendritic cells – through the release of cytokines and chemokines. They are activated through their T cell receptor in response to their specific foreign or cancer antigen, presented on MHC class II on APCs, in a multi-step process. They help to tailor the immune response depending on the specific type of pathogen encountered. Uncommitted CD4+ cells can differentiate into Th1, Th2 or Th17 cells – with distinct cytokine production patterns and functions – based on the chemical signals which they encounter.
Differentiation into Th Subtypes
The differentiation lineages of naïve CD4+ T cells are driven by a number of cytokines, transcription factors, co-stimulatory signals, and adhesion molecules, determining their final fate.
- Th1 cells: IL-12 and interferon gamma (IFNγ) are responsible for driving Th1 differentiation. IL-12 is secreted by APCs upon T cell engagement and activates the STAT4 transcription factor, promoting differentiation into Th1 effector cells. STAT4 induces IFNγ production, which further supports Th1 differentiation by activating the transcription factors STAT1 and Tbet. Tbet drives a positive feedback loop, strengthening the Th1 response through further expression of IFNγ.1,2
- Th2 cells: Differentiation into Th2 cells is specifically triggered by IL-4 and IL-2. IL-4 activates the transcription factor GATA3 through a STAT6 dependent signaling pathway.3 GATA3 interacts with IL-2-induced STAT5 signaling to upregulate expression of IL-4. This positive feedback loop drives differentiation in to Th2 cells.4
- Th17 cells: Th17 differentiation is induced by TGFβ, in combination with IL-6 or IL-21. IL-1 is also involved in early Th17 differentiation. The master regulator of differentiation is believed to be the transcription factor RORγt/RORα, although STAT3, Smads, IRF4, Ahr, and Batf are also involved.5
Th Cell Functions
- Th1: The main function of Th1 cells is the control of cell-mediated immunity and inflammation, typically against intracellular bacteria and protozoa. This occurs through the regulation of cytolytic and effector functions of other immune cells – including macrophages, B cells, and dendritic cells – triggered by the interaction of CD40 expressed on their surface with CD40 ligand (CD40L) on the surface of Th1 cells. Th1 cells secrete high levels of the cytokine IFNγ, which activates the cytolytic activities of macrophages, resulting in them killing intracellular pathogens. They also have an effect in cancer, activating cytotoxic T cells (CTLs) and NK cells to target and destroy tumor cells.6
- Th2: Th2 cells are primarily involved in the immune response against extracellular pathogens – such as parasites and bacteria – allergens, and toxins. They regulate humoral or antibody-mediated responses through the production of cytokines, including IL-4, IL-5, IL-6, IL-9, IL-13, and IL-17E (IL-25). These cytokines control phagocyte-independent responses, by regulating antibody production by B cells, eosinophil activation, and inhibition of several macrophage functions. Because of their influence on the production of antibodies and their role in allergic responses, over-activation of Th2 cells has been shown to play a role in autoimmune reactions and the aggravation of allergies.4
- Th17: Th17 are generally considered to be positive regulators of immune responses, promoting the accumulation of other immune cells at inflammatory sites mediated by IL-17A, IL-17F, IL-21, and IL-22. However, Th17 cells are very heterogeneous and display significant plasticity, with factors present in the microenvironment – including cytokines, metabolites, hormones, toxins, and nutrients – affecting Th17 cell function.7 As a result, they can also negatively regulate immune responses when necessary by secreting immunosuppressive factors, such as IL-10. Th17 cells may also contribute to pathogenic processes and have been shown to play a role in inflammation in a number of autoimmune diseases.8 In cancer, Th17 cells exhibit antitumorigenic effects through the recruitment of CD8+ T cells and IFN-γ production, but can also enhance tumor invasion and angiogenesis in some tumor types.9
Emerging Research
The diverse functions and complex interactions of the different subtypes of Th cells still raise many questions. One area of recent research has investigated the impact of genetic variations on CD4+ T cells, showing new gene associations for disease risk associated with these cells.10 The role of Th2 cells in allergy and autoimmunity, and the potential for their therapeutic modulation, is another key area of interest.11,12 While CD8+ T cell-based cancer immunotherapy is becoming well established, there is an increased understanding of the role of CD4+ T cells in the clinical response to immune checkpoint inhibition therapies.13
Tools to Study T Helper Cells
Cell Isolation
T helper cell subtypes can be isolated directly from PBMCs in whole blood using magnetic bead-based separation or FACS, directed by antibodies against specific surface markers whose expression largely corresponds to Th identity as defined by function. Alternatively, naïve CD4+ T cells can be separated using these techniques, and then activated to differentiate into an individual subtype using the associated cytokines.
Tools for Profiling Th Subtypes
Cytokine ELISAs are frequently used to profile Th subtypes, as well as to quantify the secreted cytokines. Flow cytometry is also a useful tool for profiling cells based on both cytokine expression and the presence of specific surface or intracellular markers.
Cell Markers
Activated T helper cells differentiate into functionally diverse subsets with specific cytokine expression, depending on the stimuli. Identification of T helper cells subtypes can be carried out by flow cytometry using a panel of markers.
| Th | Th2 | Th17 | ||||
|---|---|---|---|---|---|---|
| Marker type | Marker | Alternative names | Marker | Alternative names | Marker | Alternative names |
| Surface | CD4 | T4 | CD4 | T4 | CD4 | T4 |
| CD183 | CXCR3, CKR-L2, CMKAR3, GPR9, IP10-R, Mig-R, MigR, C-X-C motif chemokine receptor 3 | CD194 | CCR4, CC-CKR-4, CD194, CKR4, CMKBR4, ChemR13, HGCN:14099, K5-5, C-C motif chemokine receptor 4 | CD196 | CCR6, CKR-L3, GPRCY4 | |
| Effector cytokine | IFNƔ | IL-4 | IL-17A | CTLA8, IL-17, IL-17A, IL17, CTLA-8, , ILA17 | ||
| TNF-α | DIF, TNFA, TNFSF2, TNLG1F, | IL-5 | IL-17F | CANDF6, IL-17F, ML-1, ML1 | ||
| TNF-β | Lymphotoxin-alpha, LT-α | IL-6 | IL-21 | CVID11, Za11 | ||
| IL-13 | IL-22 | IL-D110, IL-TIF, ILTIF, TIFIL-23, TIFa, zcyto18 | ||||
| IL-26 | AK155 | |||||
| TNF-α | DIF, TNFA, TNFSF2, TNLG1F, | |||||
| CCL20 | CKb4, Exodus, LARC, MIP-3-alpha, MIP-3a, MIP3A, SCYA20, ST38, | |||||
| Transcription factor | T-bet | TBX21, T-PET, TBET, TBLYM, T-box 21, T-box transcription factor 21, IMD88 | GATA3 | HDR, HDRS, GATA binding protein 3 | RORγt | RORγ2 |
| STAT1 | CANDF7, IMD31A, IMD31B, IMD31C, ISGF-3, STAT91 | STAT5 | Signal transducer and activator of transcription 5 | RORα | RORA, NR1F1, ROR1, ROR2, ROR3, RZR-ALPHA, RZRA, IDDECA | |
| STAT6 | D12S1644, IL-4-STAT, STAT6B, STAT6C | STAT6 | D12S1644, IL-4-STAT, STAT6B, STAT6C | STAT3 | DMIO, APRF, HIES, ADMIO1 | |
Related Products
References
1. Luckheeram R, Zhou R, et al. (2012) CD4+T cells: Differentiation and functions. Clin Dev Immunol 2012:925135. doi:10.1155/2012/925135
2. Raphael I, Joern R, et al. (2020) Memory CD4+ T cells in immunity and autoimmune diseases. Cells 9(3):531. doi:10.3390/cells9030531
3. Stark J, Tibbitt C, et al. (2019) The Metabolic Requirements of Th2 Cell Differentiation. Front Immunol. 10:2318. doi:10.3389/fimmu.2019.02318
4. Walker, J, McKenzie, A (2018) TH2 cell development and function. Nat Rev Immunol 18:121–133. doi:10.1038/nri.2017.118
5. Dong C. (2011) Genetic controls of Th17 cell differentiation and plasticity. Exp Mol Med. 43(1):1-6. doi:10.3858/emm.2011.43.1.007
6. Karnell J, Rieder S, et al. (2019) Targeting the CD40-CD40L pathway in autoimmune diseases: Humoral immunity and beyond. Adv Drug Deliv Rev. 141:92-103. doi: 10.1016/j.addr.2018.12.005
7. Papadopoulou G, Xanthou G (2022). Metabolic rewiring: a new master of Th17 cell plasticity and heterogeneity. FEBS J. doi: 10.1111/febs.15853
8. Wu X, Tian J, et al (2018). Insight Into Non-Pathogenic Th17 Cells in Autoimmune Diseases. Front Immunol. 9:1112. doi: 10.3389/fimmu.2018.01112
9. Knochelmann, H, Dwyer, C, et al. (2018) When worlds collide: Th17 and Treg cells in cancer and autoimmunity. Cell Mol Immunol 15:458–469. doi:10.1038/s41423-018-0004-4
10. Schmiedel B, Gonzalez-Collins C et al. (2022) Single-cell eQTL analysis of activated T cell subsets reveals activation and cell type–dependent effects of disease-risk variants. Sci Immunol. 7(68):eabm2508. doi:10.1126/sciimmunol.abm2508
11. León B, Ballesteros-Tato A. (2021) Modulating Th2 Cell Immunity for the Treatment of Asthma. Front Immunol. 12:637948. doi: 10.3389/fimmu.2021.637948
12. Iwasaki N, Terawaki S, et al. (2021) Th2 cells and macrophages cooperatively induce allergic inflammation through histamine signaling. PLoS One. 16(3):e0248158. doi:10.1371/journal.pone.0248158
13. Lee J, Lozano-Ruiz B, et al. (2021). The Multifaceted Role of Th1, Th9, and Th17 Cells in Immune Checkpoint Inhibition Therapy. Front Immunol. 12:625667. doi:10.3389/fimmu.2021.625667

