Miniaturized Multi-Piece DNA Assembly Using the Echo 525 Liquid Handler
Jared Bailey, Jefferson Lai, Rabia Khan, and John LesnickHighlight
The Echo Liquid Handler has demonstrated the ability to generate DNA constructs using various assembly chemistries with a miniaturized protocol. Interchanging smaller, modular pieces of DNA is the preferred method for many DNA construction workflows due to decreased synthesis cost and faster progression along the design, build, test, and learn cycle. In the studies described in this Application Note, the NEB NEBuilder HiFi DNA Assembly Cloning Kit was combined with the Echo 525 Liquid Handler to assemble a construct over five modular pieces. The reaction was miniaturized forty-fold from the manufacturer recommended volume, representing a significant savings in reagent costs. After transformation into E. coli, the constructs were initially checked for successful assembly by junction colony qPCR. Then, a miniaturized Illumina MiSeq Nextera XT protocol was used for sequencing verification, allowing for further cost savings. The Echo Liquid Handler enables lower-cost methods and workflows to quickly produce high-quality synthetic DNA constructs, which expands design-based testing with higher throughput and allows the scientist to investigate a broader biological landscape.
Overview
Synthetic biology is an interdisciplinary science with the potential to impact academic and industrial applications including the creation of novel therapeutics and vaccines, plant science and biofuels. Many of these disciplines use bio-based chemical manufacturing capabilities that involve the application of engineering principles to biology. The focus is often on generating, characterizing, and isolating parts of natural biological systems before using them as components of an engineered biological pathway. A trademark of synthetic biology is the application of rational principles to the design and assembly of these biological components. Even with rational design, the impact of introducing foreign DNA into a cell can be difficult to predict. This creates the need to test multiple permutations to obtain the desired outcome. An emphasis on the modular design of DNA parts enables the assembly of a greater variety of potential constructs by interchanging the individual components. In addition to simplifying the overall DNA construction workflow, modular design and assembly of DNA components makes automation of the assembly process possible, reducing the time, labor, and cost of generating multiple constructs to allow for an increase of throughput with an overall shortened development cycle.
Synthetic DNA plasmids are designed with experimentally-dependent functionality. This can range from interrogation of different regions of a single protein to the expression of an entire biological pathway. The designed DNA plasmid is then divided into synthesizable pieces. The pieces are designed with overlapping sequences to their neighbors before being chemically synthesized. The DNA fragments are enzymatically assembled together to build the designed construct using gene assembly techniques. Multiple pieces can be assembled together into larger DNA assemblies. The assembled constructs are typically cloned into an expression vector and verified with Next-Generation Sequencing (NGS).
Once verified, the synthetic constructs are transferred into a production cell and the function of the designer construct is assayed. After learning from the assay results, the constructs can then be modified or refined. This test cycle is repeated until a DNA construct is obtained that produces the desired function.
Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) is a commonly studied gene in human oncology. The mutation of specific domains from this gene have been correlated with multiple tumor types. Due to PIK3CA’s importance in oncology, it was chosen to demonstrate a real-world application of miniaturization of the NEBuilder HiFi kit using the Echo 525 Liquid Handler. PIK3CA was divided into four modular pieces based on domains of interest and was tagged with C-terminal Green Fluorescent Protein (GFP) (Figure 1A). Four known cancer-linked mutations, R38H, E542K, E545K, and H1047R, were designed in modular pieces along with a wild-type control (Figure 1B). The construct was also engineered to assemble into a linearized vector to allow for future transfection into mammalian cells. The vector is interchangeably referred to as piece five after being linearized. Each piece was designed with 23 to 25 base pair DNA overlaps to its neighbor (Figure 1C), assembling five, five-piece assemblies by substituting modified modular DNA pieces at the nanoliter scale.
Figure 1.
A. The complete plasmid construct to be assembled with the GFP-tagged PIK3CA being inserted into the pcDNA 3.1/Zeo (+) backbone. gBlock pieces as well as the linear vector are shown in blue, with their overlapping region shown by a light green annotation labeling them as a ligation site. The PIK3CA gene is in yellow with the linker (purple) and GFP tag (green) also annotated. Other important components of the plasmid machinery are labeled as necessary.
B. Nucleotide resolution of the overlap sites that have been used to allow assembly by NEBuilder.
C. The locations of the PIK3CA functional domains (dark green) and the planned mutations (red) within the constructs. Each construct was planned as either the unmutated wild-type or with one of the four listed mutations (R38H, E542K, E545K, H1047R, or Wildtype).
Previously published work from the laboratory of Yizhi Cai, PhD at the University of Edinburgh (Kanigowska, et al. JALA 2015) has shown the ability of the Echo 550 Liquid Handler to generate two-piece assemblies using the Golden Gate and Gibson Assembly chemistries utilizing a miniaturized protocol. Additionally, we have demonstrated this five-piece reaction can be miniaturized and is robust at the 500 nL volume with 1.25 fmol of each piece as demonstrated in Table 1 in the volumes transferred by the Echo 525 Liquid Handler.
NEBuilder Reaction Conditions
| Reagent | DNA Concentration (ng/μL) | ng DNA/1.25 fmol | WT Reaction (nL) | R38H Reaction (nL) | E542K Reaction (nL) | E545K Reaction (nL) | H1047R Reaction (nL) | Echo Calibration |
| Piece 1 WT | 23.6 | 16 | 25 | 25 | 25 | 25 | 384LDV_PIus_AQ_GP |
|
| Piece 1 R38H | 25 | 16 | 25 | |||||
| Piece 2 WT | 14.6 | 11 | 50 | 50 | 50 | 50 | 50 | |
| Piece 3 WT | 7.69 | 14 | 75 | 75 | 75 | |||
| Piece 3 E542K | 10 | 14 | 75 | |||||
| Piece 3 E545K | 10 | 14 | 75 | |||||
| Piece 4 WT | 29.6 | 26 | 50 | 50 | 50 | 50 | ||
| Piece 4 H1047R | 30 | 26 | 50 | |||||
| Piece 5 | 83.9 | 83 | 50 | 50 | 50 | 50 | 50 | |
| NEBuilder Master Mix | 250 | 250 | 250 | 250 | 250 |
Table 1. Volumes of reagents used for the modular DNA assemblies. The five desired assembly reactions are listed as headers with the requisite volume for each piece listed below in nanoliters. All of the reactions had approximately 1.25 fmols of each piece of insert DNA added. Finally, 250 nL of the NEBuilder Master Mix was added to bring the final volumes to 500 nL.
The modular DNA assemblies were transformed into chemically competent NEB 10B E. coli. The Echo 525 Liquid Handler was then used to array 8 columns of 25 nL per construct transformation onto OmniTrays using carbenicillin as selection. The resulting isolated colonies plates were checked for quality control by colony qPCR and then sequenced using NGS. The Echo 525 Liquid Handler was used to accurately dispense a cell suspension into four qPCR reactions targeting different overlap junction regions of the assembled plasmid construct.
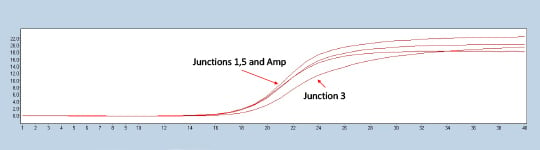
Figure 2. Lightcycler 480 trace of a representative construct testing positive for all three experimental junctions in comparison to the ampicillin marker control.
Colonies with all three junction Crossing points (CPs) closely matched to the ampicillin control were called positive assemblies (Figure 2). Closely matching the CP of a known control to the internal junctions allows the scientist to check whether the completed plasmid is present in the expected orientation and ratio. The sequencing of the validated plasmids can be done in a cost-effective way using a miniaturized Nextera XT protocol in conjunction with the Illumina MiSeq. Prior in-house work has demonstrated the ability to miniaturize the Nextera XT kit ten to one hundred-fold. In the Application Note Effective Miniaturization of Illumina Nextera XT Library Prep for Multiplexed Whole Genome Sequencing and Microbiome Applications, ten-fold Nextera XT reaction volumes were confirmed to also work for plasmid validation as seen in Figure 3.
Increasing the throughput of DNA construction is a goal of many research groups. The use of modular DNA pieces within a construction pipeline is a standard practice that offers the benefit of both lower synthesis cost when compared to purchasing a full-length gene, and faster lead consolidation upon successful screening. The Echo 525 Liquid Handler, in combination with the NEBuilder HiFi kit, can further enable this technique through the miniaturization of the reactions and workflow enhancements. The use of a miniaturized 500 nL reaction volume while varying modular component pieces for a desired mutation was the primary aim of this experiment. The NEBuilder HiFi kit reaction volume was decreased forty-fold from the recommended volume, from 20μL (20,000 nL) down to 500 nL. At the 500 nL reduced volume, only 1.25 fmoles of DNA was needed for each input piece, which dramatically reduced the amount of DNA required from that of the recommended protocol of 500 fmoles per piece.
Obtaining sufficient, high-quality input DNA is challenging at a high-throughput scale, so the decrease of the required number of DNA molecules can vastly increase the number of modular assemblies possible in a building campaign. These smaller reagent volumes of PIK3CA constructs containing the mutations identified as R38H, E542K, E545K, and H1047R were used, as well as wild-type, in a mammalian expression vector. Correctly assembled clones were obtained for each of these targets as seen in the alignment from Figure 3. Successfully mutated PIK3CA constructs generated through the acoustic arraying of interchangeable modular pieces at nanoliter volumes allows for a large scale increased-throughput DNA assembly workflow due to reduced reaction cost and DNA requirements.
Several other process improvements used in this study would help with an increase in the DNA assembly pipeline at larger scales. One such process improvement demonstrated in this experiment is the ability to acoustically plate the transformants onto solid media. OmniTray plating with the Echo 525 Liquid Handler can save operator time as well as incubator space compared to traditional transformation plating. The Echo 525 Liquid Handler is uniquely positioned to make this plating possible. Tip-based systems have difficulty with poured agar plates being inherently concave and variable in height. The Echo 525 Liquid Handler transfers 25 nL increments of cell suspension very precisely onto the solid media on an X, Y grid, but with a buffered Z height due to the acoustic, tipless transfer of the droplet itself. This allows the transfer of cells onto solid media with minimal impact from the media height.
Another major process improvement was seen using colony qPCR for the transformed NEBuilder reactions. One of the major difficulties with any DNA construction technique is the removal of false positives, which can exceed 50% of clones after sequence verification. Background vector is especially problematic due to its ability to replicate and allow the E. coli to grow on the antibiotic selection used. Using colony qPCR on the Echo 525 Liquid Handler, the background colonies can be removed prior to the time-consuming plasmid recovery step. Colony qPCR increases the number of screened clones possible when compared to end-point colony PCR due to the removal of the laborious, low-throughput use of agarose gels. The quantitative results provided by the colony qPCR also remove the subjective nature of agarose gel reading. The colony qPCR method was able to correctly predict 100% of vector background present, as well as 74% of the insertion deletion constructs from 192 samples sequenced allowing for a vastly enriched sequencing sample set.
Finally, the sequencing of the passed plasmids can be done in a cost-effective way using a miniaturized Nextera XT protocol in conjunction with the Illumina MiSeq. Prior work had demonstrated the ability to miniaturize the Nextera XT kit ten to one hundred-fold. In this experiment, ten-fold Nextera XT reaction volumes were confirmed to also work for plasmid validation, allowing for a reduction of overall reagent needs for NGS plasmid validation. Beyond overall DNA assembly reaction volume reduction, use of the Echo 525 Liquid Handler dramatically improves the DNA construction pipeline.
Working at the nanoliter scale in synthetic biology would have been inconceivable in the recent past, but the Echo Acoustic Technology makes these volumes a reality. Complex, modular DNA assemblies were generated using the NEBuilder HiFi kit at a forty-fold reduction in volume, significantly reducing costs and allowing for substantial increases in throughput. The integration of a Echo 525 Liquid Handler into a DNA construction pipeline, such as one using the NEB NEBuilder HiFi kit, can allow for significant volume reductions as well as workflow enhancements with both complex and simple plasmid designs.
Figure 3. Illumina MiSeq reads from each of the five constructs aligned against the wild-type (unmutated) PIK3CA gene. Sequence coverage and location of the determined mutation are shown in the consensus bar at the top of each window. A magnified region containing the mutated amino acid residues are shown next to the four mutated versions of PIK3CA.
This protocol is for demonstration only, and is not validated by Beckman Coulter. Beckman Coulter makes no warranties of any kind whatsoever express or implied, with respect to this protocol, including but not limited to warranties of fitness for a particular purpose or merchantablity or that the protocol is non-infringing. All warranties are expressly disclaimed. Your use of the method is solely at your own risk, without recourse to Beckman Coulter. Not intended or validated for use in the diagnosis of disease or other conditions.
Beckman Coulter Life Sciences liquid handlers are not intended or validated for use in the diagnosis of disease or other conditions.