Biomek i-Series Automated KAPA HyperPrep and HyperPlus Workflows
Introduction
The KAPA HyperPrep and HyperPlus Kits allow users to prepare DNA libraries compatible with Illumina sequencing platforms from a variety of sample types (including high-quality genomic DNA, FFPE derived DNA, etc.) and a wide range of input amounts (1 ng to 1 µg). The KAPA HyperPrep Kit utilizes DNA that has been acoustically sheared utilizing a Covaris or equivalent system, while the KAPA HyperPlus Kit utilizes a proprietary enzymatic fragmentation system to fragment DNA prior to downstream library preparation using KAPA HyperPrep chemistry. Both kits are compatible with dual or single-indexing strategies and can process up to 96 samples per kit.
In comparison to manual pipetting, automation of these workflows on Biomek platforms provides:
- Reduced hands-on time and increased throughput;
- Reduction in potential pipetting errors;
- Standardized workflow for improved results;
- Quick implementation with ready-to-install methods; and
- Knowledgeable support.
In this flyer, the automated performance of KAPA HyperPlus Kit on the Biomek i7 Hybrid Genomics Workstation is demonstrated. Data showing the performance of the KAPA HyperPrep Kit on the Biomek i7 Hybrid Genomics Workstation was collected but is not shown.
Automated method
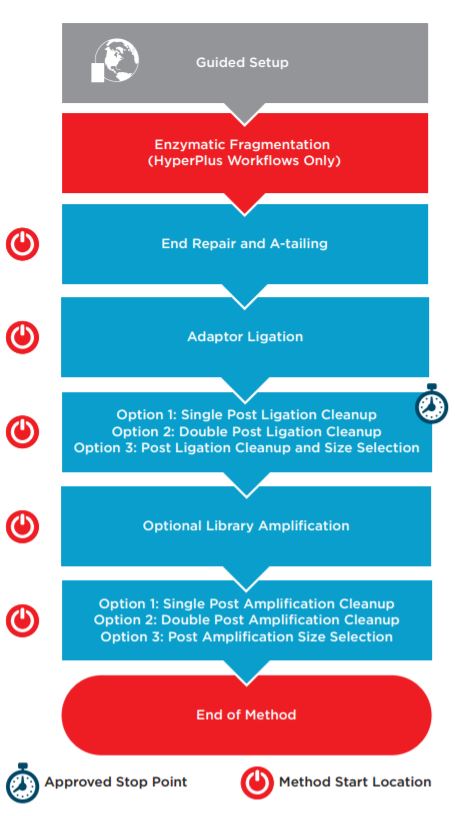
Figure 1. KAPA HyperPrep/HyperPlus automated method workflow.
Automation provides increased efficiency by reducing hands-on time (Table 1). The complete method (Figure 1) can be run with full walk away capability but does include logical start and stop points assigned based on Roche’s recommendations. This provides users flexibility in workflow scheduling, allowing each laboratory to address their individual requirements for sample processing and throughput. Each section of the workflow shown above with a Method Start Location icon can serve as a place to begin for error-recovery purposes. The automation method also includes a variety of options for post-ligation and post-amplification SPRI (Solid Phase Reversible Immobilization) cleanups using AMPure XP, allowing the user to select a single cleanup, a double cleanup, or a single cleanup followed by size selection.
| Metric | KAPA HyperPrep | KAPA HyperPrep PCR Free | KAPA HyperPlus | KAPA HyperPlus PCR Free |
| Sample Throughput | 96 | 96 | 96 | 96 |
| Hands On Time | 20 min | 20 min | 20 min | 20 min |
| Biomek Run Time | 3 hr, 50 min | 2 hr, 41 min | 4 hr, 2 min | 2 hr, 52 min |
| Total Run Time | 4 hr, 10 min | 3 hr, 1 min | 4 hr, 22 min | 3 hr, 12 min |
| Number of User Interactions (Aside From Initial Setup) | 0 | 0 | 0 | 0 |
Table 1. Estimated run times for the KAPA HyperPrep and HyperPlus Kits on the Biomek i7 Hybrid Genomics Workstation. Timing estimates based a single post-ligation cleanup, and if PCR is conducted, 10 cycles of library amplification and a single post-amplification cleanup.
The software provides several user-friendly features such as:
1. Biomek Method Launcher (BML)
BML is a secure interface for method implementation without affecting method integrity. It allows the users to remotely monitor the progress of the run. The manual control options provide the opportunity to interact with the instrument, if needed.
Figure 2. Biomek Method Launcher provides an easy interface to launch the method.
2. Method Options Selector (MOS)
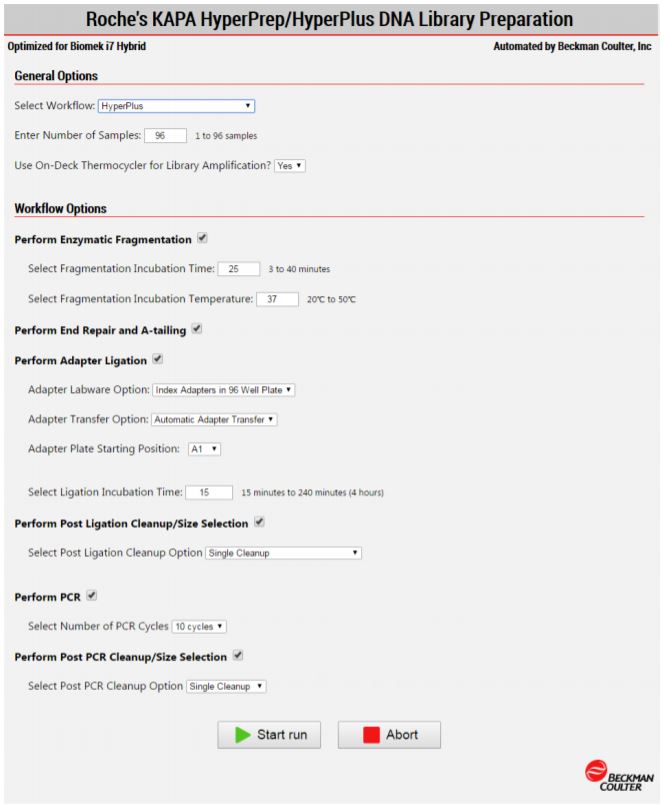
MOS enables selection of plate processing and sample number options to maximize flexibility, adaptability and the ease of method execution.
Figure 3. Biomek Method Options Selector enables users to select the desired workflow, sample number, and a variety of workflow customization options.
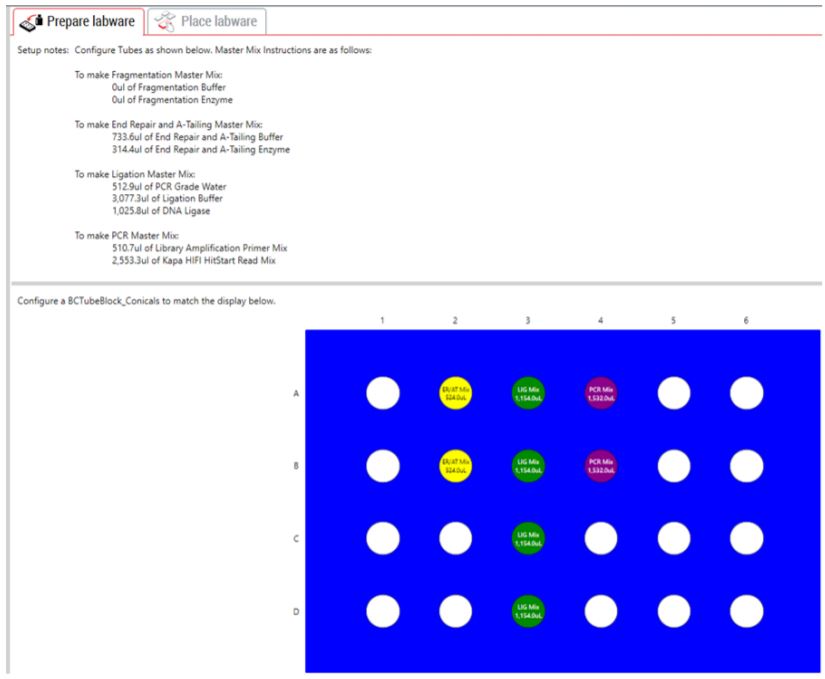
GLS is generated based on options selected in the MOS and provides the user specific graphical setup instructions with reagent volume calculation and step-by-step instructions to prepare reagents.
Figure 4. Guided Labware Setup indicates reagent volumes and guides the user for correct deck setup.
Experimental Design
In Experiment 1, Coriell NA12891 DNA was used as template for a KAPA HyperPlus run of eight samples, testing the upper and lower limits of the workflow, along with a midpoint. Three 1 ng replicates, three 50 ng replicates and two 1000 ng replicates were produced, along with a set of manually prepared libraries for comparison. DNA was fragmented for 30 minutes at 37ºC. The manual libraries were prepared according to Roche specifications, while the automated libraries were prepared using more stringent post-ligation and post-amplification cleanups (0.75X and 0.95X SPRI ratios, respectively). Library amplification was performed and 1 ng, 50 ng, and 1000 ng inputs were cycled separately (16, 6, and 3 cycles, respectively). For libraries produced on the Biomek i7, all incubations and library amplification were performed on deck using the Life Technologies Automated ThermoCycler (ATC). Library size distributions were assayed on the Agilent 2100 Bioanalyzer using the High Sensitivity DNA assay. Post-ligation and post-amplification yields were assessed using the KAPA Library Quantification Kit.
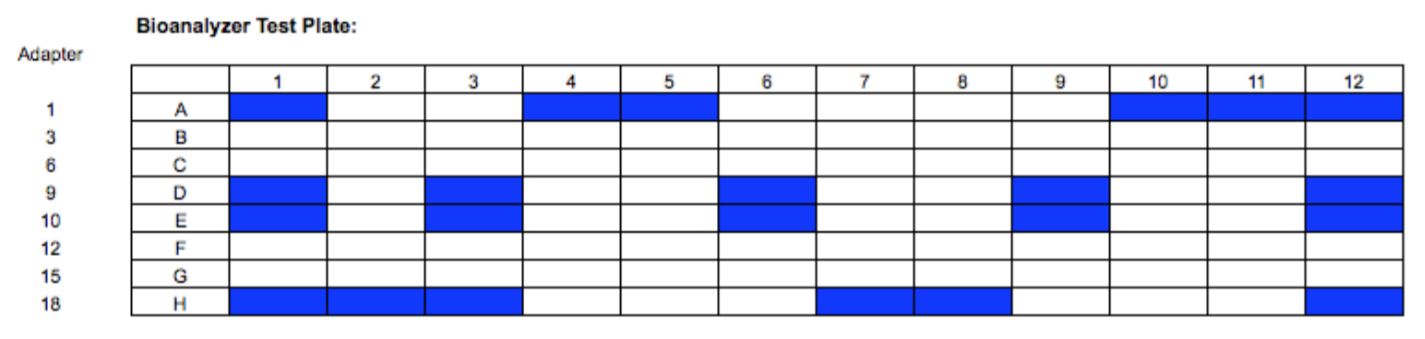
In Experiment 2, a full 96 well plate was processed to assay for the presence of plate effects. A single DNA input amount was used (50 ng Coriell NA12891), and all library preparation and automation conditions were kept consistent with Experiment 1. Final library concentration for all 96 samples was measured using Qubit. Library size distribution was assessed for a subset of libraries (Figure 5) using the High Sensitivity DNA assay on the Agilent 2100 Bioanalyzer.
Figure 5. Visual representation of which libraries were selected for sizing assessment.
Experimental Results and Discussion
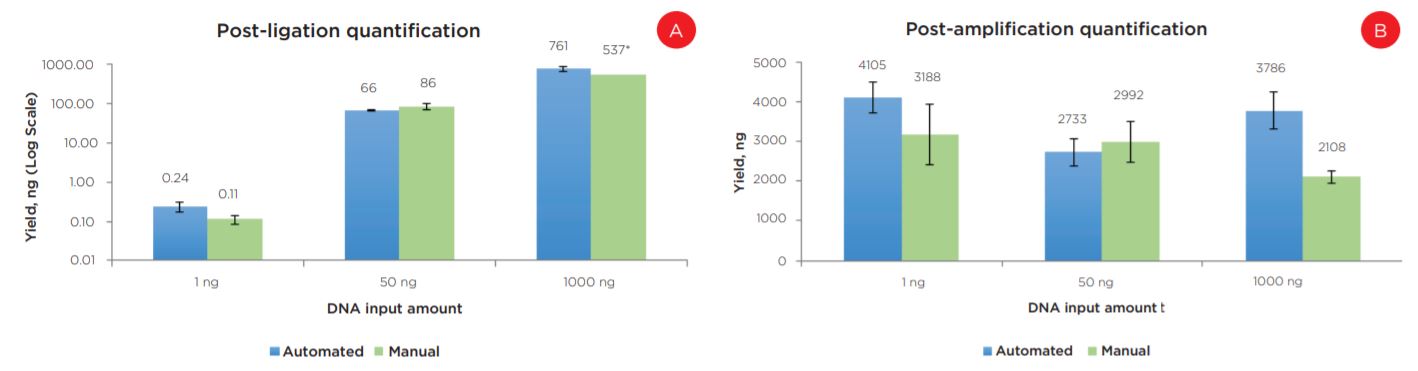
Experiment 1 results showed equivalent post-ligation and post-amplification yields between the automation and manually prepared libraries across a wide range of input amounts (Figures 6a and 6b). This suggests equivalency with respect to library complexity, as well as amplification efficiency. Additionally, library size distributions were comparable between automated and manual preparations. Representative data is shown for 50 ng libraries (Figure 7).
Figure 6. Post-ligation (a) and post-amplification (b) yields indicate the automated method produces libraries with comparable yields to manually prepared samples.
* Due to a qPCR error, data is shown for a single replicate of this condition.
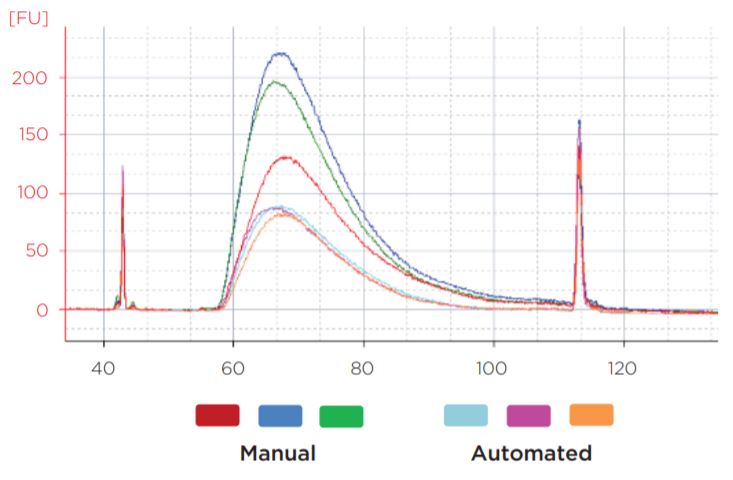
Figure 7. Representative library traces of the 50 ng samples show consistent sizing between automated and manual libraries. The average mode size of the manual libraries was 267 bp, while the automated library average mode size was 266 bp.
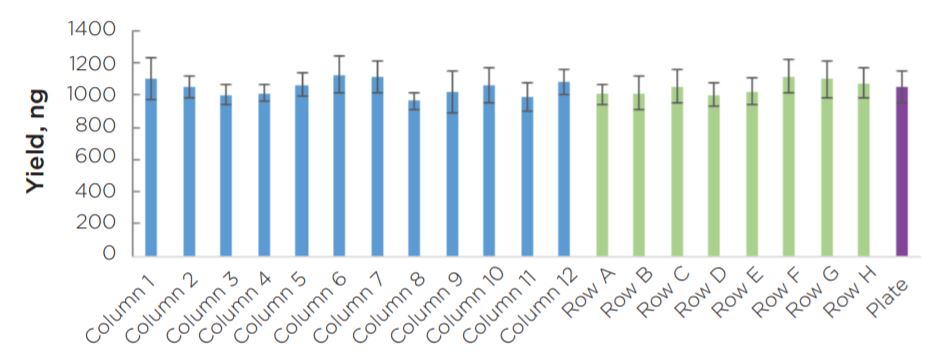
Experiment 2 final library yields showed consistent results across the plate (Figure 8). All individual columns and rows show average yields that are comparable to each other, as well as to the full plate average. Similar to the yield data, sizing results showed consistency across individual columns and rows in comparison to the full plate average mode (Figure 9). Overall, results do not indicate the presence of significant plate effects.
Final Library Quantification
Figure 8. Final library yield of all 96 KAPA HyperPlus libraries generated by the automated method are consistent across columns and rows.
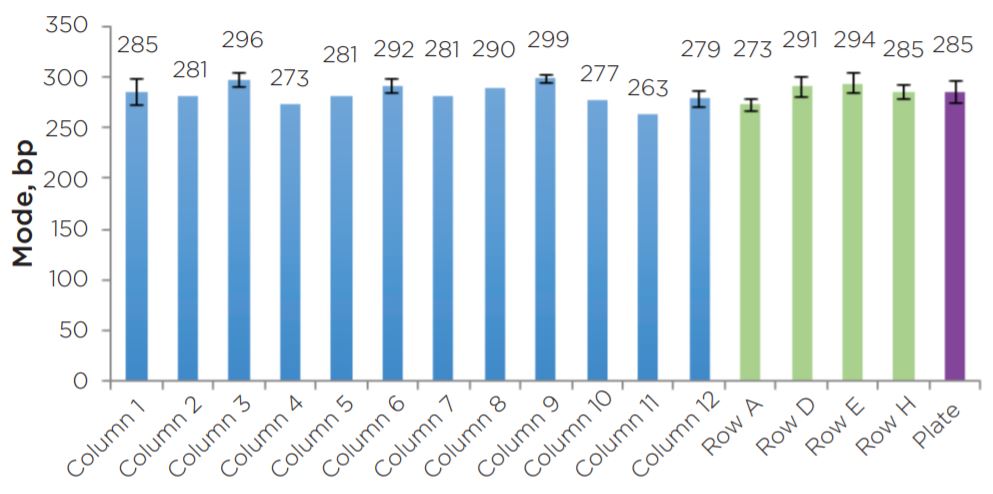
Average Mode
Figure 9. The average modes of the 22 libraries assayed show consistent sizing across columns and rows.
Summary
A robust and reliable automation solution for NGS library prep is essential to take full advantage of Illumina’s sequencing technology. Automation saves valuable time and money, giving researchers an efficient process with more walk-away time.
Performance of the KAPA HyperPlus workflow on the Biomek i7 Hybrid Genomics Workstation has been demonstrated, and results show that the method delivers high-quality libraries on par with those prepared manually. The solution is fully walk-away and scalable, thus providing fast and efficient library construction.