Build Analysis Workflows by Combining the Power of Kaluza Analysis Software and the Cytobank Platform
Machine Learning approaches have become a necessity for the in-depth analysis of complex immunophenotyping data and the unbiased evaluation of groups of samples and conditions. Researchers often need to learn new tools and manually combine results from different platforms to generate insights.
Using the Kaluza Cytobank Plugin* allows the user to move compensated, transformed and anonymized data sets from the Kaluza desktop software to the Cytobank cloud-platform*, taking advantage of the intuitive compensation adjustments supported by Kaluza Analysis Software* and the machine-learning algorithms integrated in the Cytobank platform. Combining both packages offers the user the flexibility to choose from a broad offering of features and functionalities and to customize their analysis pipeline to fit their research needs. Used in combination, Kaluza and the Cytobank platform offer an easy to use end to end data analysis and management solution.

Download the Kaluza Cytobank Plugin from the Kaluza Downloads page.
For step by step instructions refer to Leveraging the Combined Power of Kaluza and the Cytobank Platform in the Learning Center.
The Kaluza Cytobank plugin enables the combination of Kaluza features that allow intuitive scale adjustments, easy compensation corrections and the fast development of a gating strategy with the advanced algorithms and data management offered by the Cytobank platform.
Different data visualization tools support the exploration of low to medium complexity data. In addition to real time data exploration tools Kaluza Analysis offers instrument and assay QC tracking with a dedicated module. Data reporting and batch analysis functionalities enable the rapid generation of summary results.
The Cytobank software is a cloud-based analysis platform with integrated machine-learning based analysis algorithms, as well as a structured and secure content management system for flow cytometry and other single cell data.
Cytobank’s clustering, dimensionality reduction, and visualization tools (SPADE, viSNE, CITRUS, FlowSOM) leverage the scalable compute and collaborative power of the cloud, allowing large analyses to be done quickly, and the cloud-based storage provides the capability to automatically archive and easily share these data securely and safely1.
Analysis of multi-dimensional flow cytometry data is done in several stages2. Early stages of data QC and cleanup have lower requirements for compute performance and may be performed on your own laptop or PC. Compensation adjustment, data transformation and subset identification are then followed by exploration and clustering.
The exploration stage provides an overview of all cells in the samples. For low to mid-complexity marker panels, bi-parametric plots supplemented with tools such as Radar Plots and Tree Plots can be sufficient. For high dimensional data, dimensionality reduction techniques such as viSNE allow the information from all parameters to be summarized in a single two-dimensional display4.
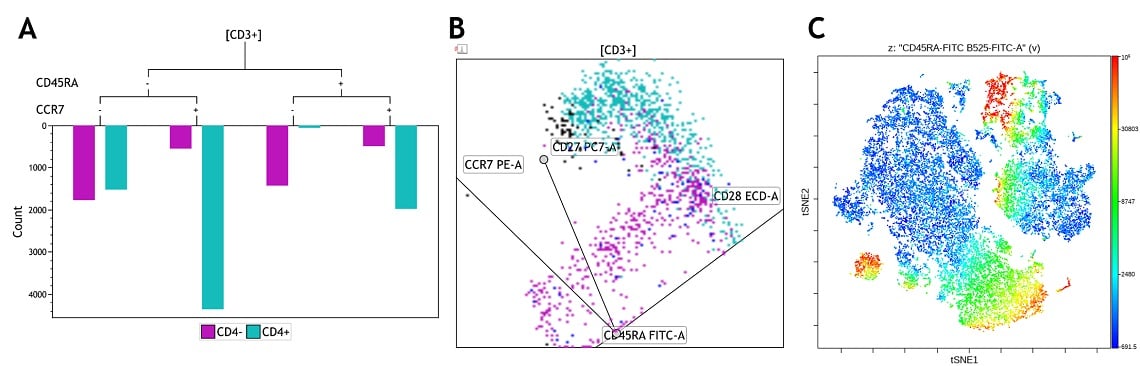
Different visualizations of T cell memory marker expression on CD4+ and CD8+ T cell subsets. Plots are for demonstration purposes only, Kaluza Tree Plot (panel A), Kaluza Radar Plot (panel B) and Cytobank viSNE (panel C). Data was generated using a normal whole blood sample stained with DURAClone IM T Cell Subsets Tube * (Part Number B53328), acquired on a CytoFLEX LX cytometer * (Part Number C40324) and analyzed using Kaluza Analysis Software * and the Cytobank Platform *.
In the next stage of data analysis, clustering algorithms such as FlowSOM and SPADE can be used to automatically identify cells of similar phenotype in an unbiased manner 5,6.
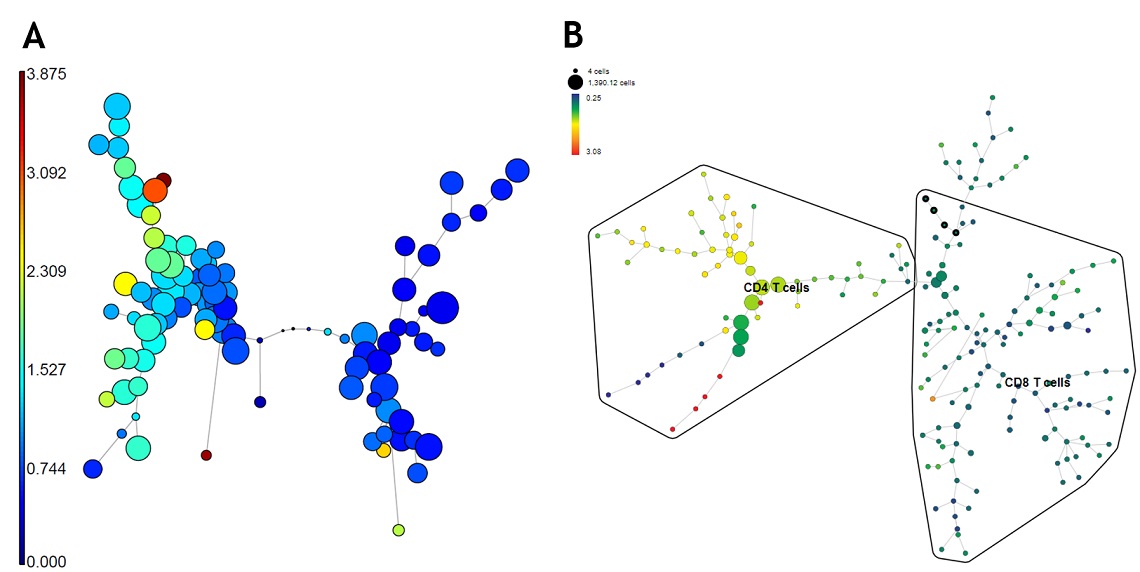
Shown are different of clustering methods based on T-cell and T cell memory marker expression. Plots are for demonstration purposes only. Cytobank FlowSOM channel colored Minimum Spanning Tree (panel A), Cytobank SPADE Minimum Spanning Tree (panel B). Data was generated using a normal whole blood sample stained with DURAClone IM T Cell Subsets Tube* (Part Number B53328), acquired on a CytoFLEX LX cytometer * (Part Number C40324) and analyzed using the Cytobank Platform*.
These later analysis stages benefit from cloud computing to free up local equipment and allow for parallel algorithm runs. While it is possible to perform viSNE, SPADE and FlowSOM calculations using desktop software, the computer cannot be used for other tasks during this time. Also, only one run at a time can be performed, slowing down optimization of algorithm settings. Cloud-based data analysis does not suffer from these limitations.
In order to combine the comfort of desktop based real time data interaction with powerful cloud-based machine learning tools, a solution for smooth data transfer is required. The Kaluza Cytobank Plugin bridges this gap by enabling researchers to compensate and transform their data using Kaluza‘s innovative user interface and real time feedback and directly upload the data to their Cytobank account.
References
- Kotecha, N., Krutzik, P. O. & Irish, J. M. Web-Based Analysis and Publication of Flow Cytometry Experiments. Current Protocols in Cytometry 2010:53, 10.17.1-10.17.24.
- Rahim, A. et al. High throughput automated analysis of big flow cytometry data. Methods 2018:134–135, 164–176.
- Mair, F. et al. The end of gating? An introduction to automated analysis of high dimensional cytometry data: Highlights. European Journal of Immunology 2016:46, 34–43.
- Amir, E. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nature Biotechnology 2013:31, 545–552.
- Qiu, P. et al. Extracting a cellular hierarchy from high-dimensional cytometry data with SPADE. Nature Biotechnology 2011:29, 886–891.
- Van Gassen, S. et al. FlowSOM: Using self-organizing maps for visualization and interpretation of cytometry data: FlowSOM. Cytometry 2015:87, 636–645.
* For Research Use Only. Not for use in diagnostic procedures.

